Having breakfast at the end of a conference is a good way to recap what was discussed and can help to generate new ideas. Two years ago at ScienceOnline09 a conversation with Cameron Neylon that followed up on the session Reputation, authority and incentives. Or: How to get rid of the Impact Factor (moderated by Björn Brembs and Pete Binfield) was started my interest in unique author identifiers for researchers. An interview with Geoff Bilder about author identifiers and the CrossRef Contributor ID project followed a month later – still one of my favorite blog posts. I have since become deeply involved with author identifiers and have joined the Board of the Open Researcher & Contributor ID (ORCID) initiative last September.
Wednesday to Friday I attended the Beyond the PDF workshop in San Diego to discuss how we can do better in scholarly publishing. The limitations of the PDF format were just one topic, the main themes were annotation, data, provenance, new models, writing and reviewing and impact. This is my presentation:
We had two very productive breakout sessions about writing and reading tools, and we agreed that we should build something that makes it much easier to describe and distribute our research data. Most people in the group took a very pragmatic approach and want to build simple tools appropriate for small research groups in the next few months. We thought that graduate students would be good early adopters, and we already have three principal investigators willing to test these tools in their labs.
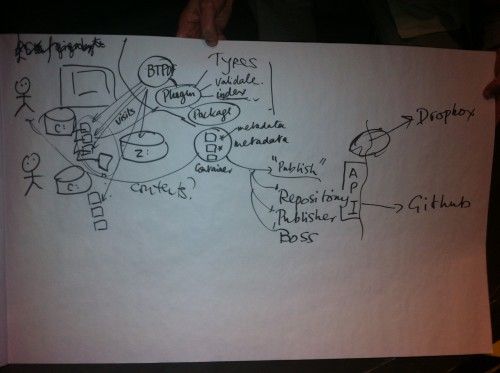
Peter Sefton demonstrated his Fascinator tool that already has a lot of the required functionality. But it was the breakfast discussion before departing – again with Cameron Neylon, but this time also including Peter Murray Rust, Peter Sefton and Ana Nelson – that helped me to put all my thoughts into place.
ePub should become the standard document format for authoring, distributing and reading scholarly content.
The ePub format uses a collection of files held together in a zip archive. Content is displayed using a combination of XHTML and CSS – not different from web pages – and the ePub can also contain other files. Journal publishers use XML internally, and it is therefore easy to distribute journal articles in ePub format – some of them are already doing this routinely. ePub has several advantages over PDF, including:
- ePub can be used for all steps in the creation of a scholarly document, including data collection, authoring, annotating and peer review. There is no need for time-consuming and expensive format conversions. Currently most manuscripts are submitted in Microsoft Word or LateX formats, and then converted first to XML and then to HTML and PDF. Metadata such as author identifiers, digital object identifiers and semantic information can be added early on and don’t get lost in a format conversion.
- ePub makes it easy to include supplementary material, e.g. video and other multimedia content, the datasets used in the publication (particularly the data used for tables and figures), all cited references in BibTeX format, etc.
- ePub is much better suited for reading on mobile devices, as the format allows reflowing of content. Most articles today are printed from the PDF and then read, but this behavior is rapidly changing.
ePub is relatively new, and not many applications for scientists already support this format. We want lab equipment that stores its data in ePub, lab notebooks that write all files from an experiment into an ePub file, reference managers that store and display papers in ePub format, authoring tools that import all these ePub files and thus make it much easier to write, annotate and submit a manuscript, and journal submission systems that take ePub files. I have written a lot about WordPress recently and this is of course a platform that would play nicely with ePub. At least two WordPress plugins support ePub, and it should be possible to modify them to the requirements of the scholarly paper.
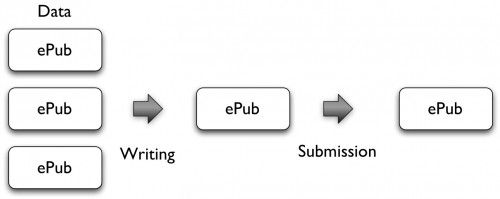
Almost as important as the document format is the distribution mechanism of these ePub files. We need a system that makes it easy to collaborate on a document, and that includes version control. The simplest solution would of course be centralized and web-based, but I’m not sure that this is a realistic scenario. We talked a lot about Dropbox during the meeting, but a solution using git (and github), Amazon Simple Storage Service, Windows Live SkyDrive or the repository software ePrints or DSpace is also possible. As an ePub document can contain all required documents, the submission of a manuscript to a journal or institutional repository could become as simple as uploading a single file, and all the peer review (including reviewer comments and revisions) could be done with that file. Submissions of datasets to databases such as Dryad could of course also be done using ePub files. A versioned distribution system should also make it easier to automatically get information about corrections or retractions (e.g. using the CrossMark system that will launch in 2011) and to receive regular updates of article-level metrics, including new citations of the article or dataset.
Several of us attending the meeting will continue the discussion in the coming weeks, and I hope I can convince them of the advantages of ePub. It shouldn’t take us more than a month or two to produce a nice ePub of the sample PLoS Computational Biology article provided for the Beyond the PDF workshop. The next Science Online London Conference will be September 2-3 at the British Library. This is a good opportunity to discuss the progress of this project, ideally including reports about new ePub tools for scientists, more journals using ePub for their articles, and practical feedback from the first users.


